Gallard A, Foucras G, Coureau C, Guéry J-C (2002) Tracking T cell clonotypes in complex T lymphocyte populations by real-time quantitative PCR using fluorogenic complementarity-determining region-3-specific probes.Costello M, Fleharty M, Abreu J et al (2018) Characterization and remediation of sample index swaps by non-redundant dual indexing on massively parallel sequencing platforms.Best K, Oakes T, Heather JM et al (2015) Computational analysis of stochastic heterogeneity in PCR amplification efficiency revealed by single molecule barcoding.Mamedov IZ, Britanova OV, Zvyagin IV et al (2013) Preparing unbiased T-cell receptor and antibody cDNA libraries for the deep next generation sequencing profiling.Shugay M, Britanova OV, Merzlyak EM et al (2014) Towards error-free profiling of immune repertoires.Weinstein JA, Jiang N, White RA et al (2009) High-throughput sequencing of the zebrafish antibody repertoire.Peacock T, Heather JM, Ronel T, Chain B (2021) Decombinator V4: an improved AIRR compliant-software package for T-cell receptor sequence annotation? Bioinformatics (Oxford, England) 37:876–878.Thomas N, Heather J, Ndifon W et al (2013) Decombinator: a tool for fast, efficient gene assignment in T-cell receptor sequences using a finite state machine.Oakes T, Heather JM, Best K et al (2017) Quantitative characterization of the T cell receptor repertoire of Naïve and memory subsets using an integrated experimental and computational pipeline which is robust, economical, and versatile.Uddin I, Woolston A, Peacock T et al (2019) Quantitative analysis of the T cell receptor repertoire. 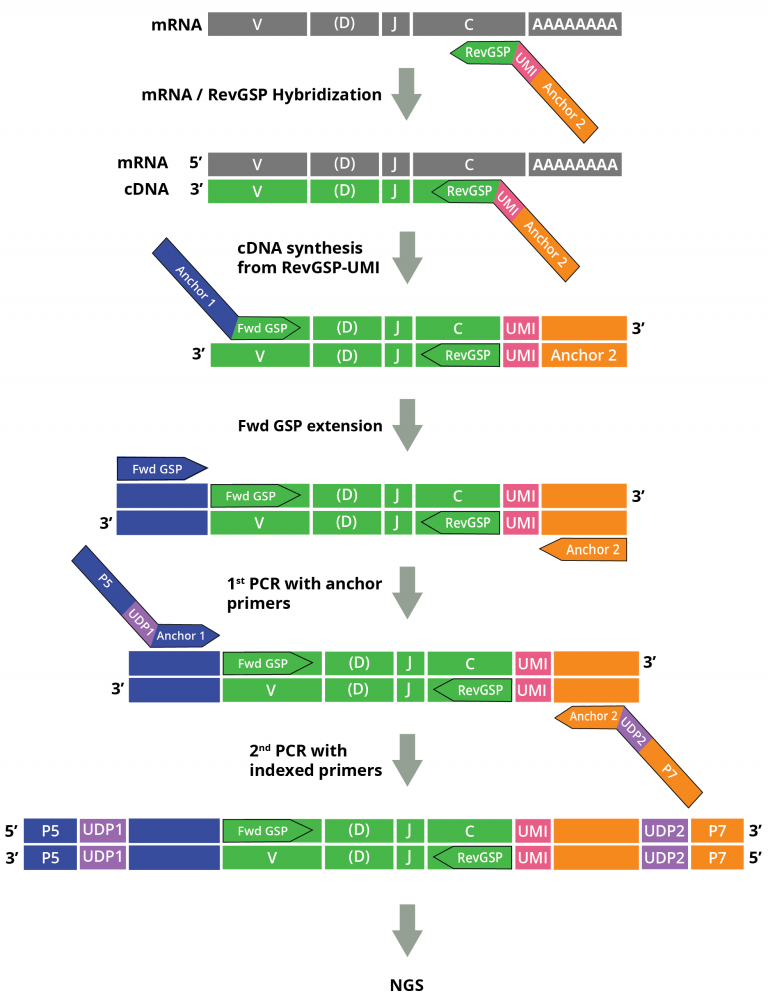
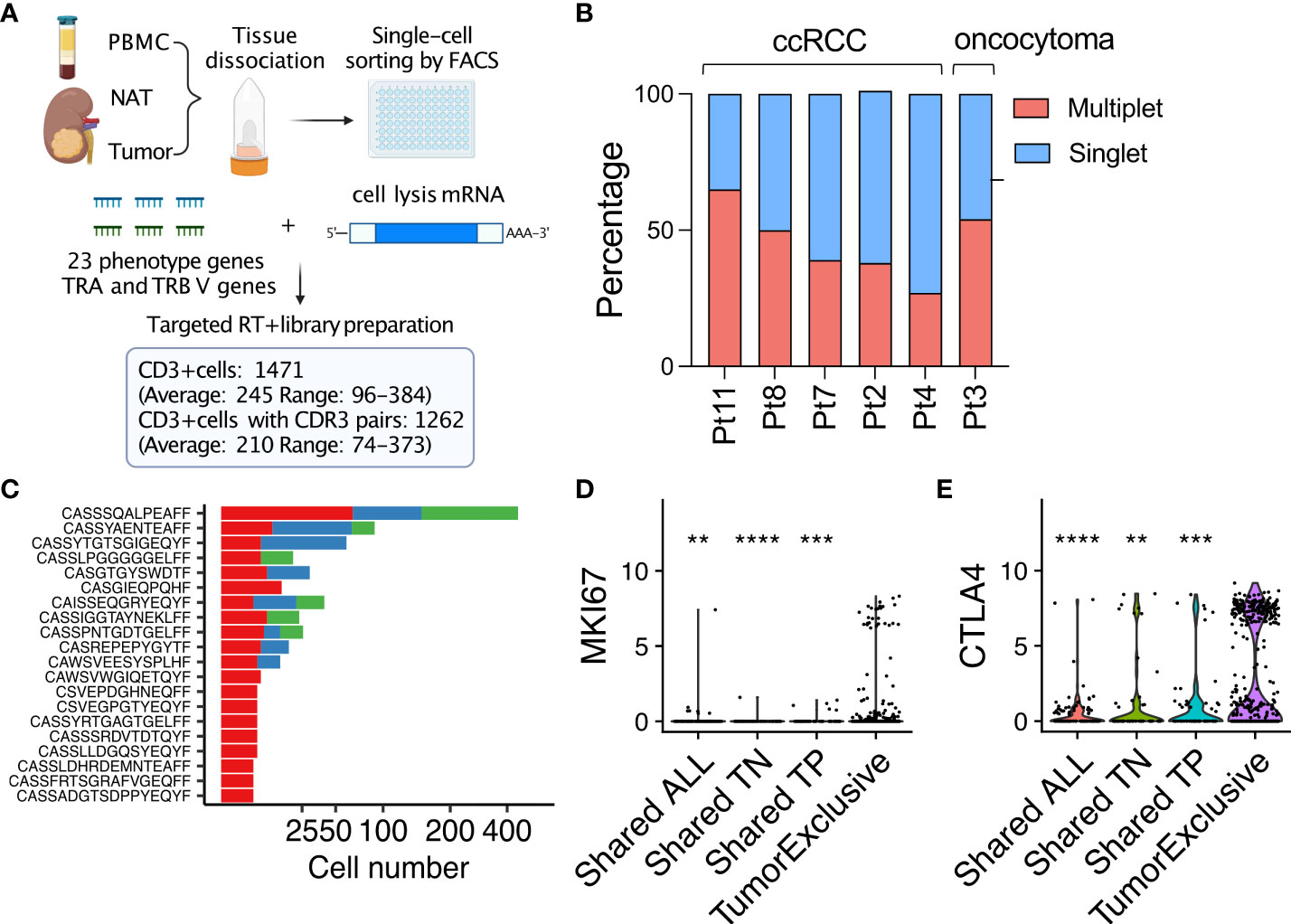
Barennes P, Quiniou V, Shugay M et al (2021) Benchmarking of T cell receptor repertoire profiling methods reveals large systematic biases.Heather JM, Chain B (2016) The sequence of sequencers: the history of sequencing DNA.Joshi K, Milighetti M, Chain BM (2021) Application of T cell receptor (TCR) repertoire analysis for the advancement of cancer immunotherapy.This method has been applied to the analysis of tumor-infiltrating lymphocytes and matched peripheral blood samples from patients with a variety of solid tumors. Samples are then tagged with unique pairs of indices, facilitating robotic scale-up and significantly reducing cross-sample contamination from index hopping. After sequencing, this molecular identifier can be used to correct both sequence errors and the effects of differential PCR amplification efficiency, thus producing a more accurate measure of the true T cell receptor frequency within the sample. The key experimental step is the ligation of a single-stranded oligonucleotide to the 3′ end of the T cell receptor cDNA, which allows easy amplification of all possible rearrangements using only a single set of primers per locus, while simultaneously introducing a unique molecular identifier to label each starting cDNA molecule. We describe an open-source protocol for amplifying, sequencing, and analyzing T cell receptors which is economical, robust, sensitive, and versatile. Most published data on the T cell repertoire continue to rely on commercial proprietary methods, which often do not allow access to the raw data, and are difficult to validate. Adaptive immunity recognizes and responds to tumors, although they are part of the immunological “self.” T cells, both CD4+ and CD8+, play a key role in the process, and the specific set of receptors which recognize tumor antigens therefore has the potential to provide prognostic biomarkers for tracking tumor growth after cancer therapy, including immunotherapy.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
